CRISPR - CAS9 GENOME EDITING
CRISPR OVERVIEW
This system constitutes a simpler and faster alternative to genome editing technologies such as ZFNs and TALENs.
It is based on the bacterial clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 system which has been adapted to serve as a versatile platform for RNA-directed genome editing in mammalian cells.
It requires three different elements:
- CRISPR RNAs (crRNAs)- a 20 BPs sequence complementary to the target sequence in the genome.
- trans-activating crRNA (tracrRNA)
- CRISPR-associated (Cas) proteins
Target cleavage by Cas9 requires base-pairing between the crRNA and tracrRNA (which can also be combined in a single hybrid guide RNA) as well as base pairing between the crRNA and the target DNA. Target recognition is facilitated by the presence of a short sequence called a protospacer-adjacent motif (PAM) that conforms to the sequence NGG, located in the complementary strand of that recognized by crRNA.
These events result in the occurrence of DNA Double Strands Breacks (DSBs). These can be repaired by two different mechanisms occurring in the cell:
- non-homologous end joining (NHEJ) and
- homology-directed repair (HDR),
These two different break repair pathways can be used respectively to generate gene knock-outs or knock-in of defined modifications at the site of cleavage. NHEJ is an error-prone process that frequently results in the formation of small insertions and deletions that disrupt gene function. HDR requires homologous DNA as a template (donor DNA) flanked on either side by sequences bearing homology to the target.
The simplicity of using of a Using the CRISPR/Cas9 strategy, retargeting the nuclease complex only requires introduction of a new RNA sequence and there is no need to reengineer the specificity of protein transcription factors.
CRISPR/CAS9 METHODS
Both CAS9 and guide RNA can be decoded by plasmids or, conversely, CAS 9 can be introduce in the cell as DNA expressed by a vector and guide RNA can be synthesized as an oligonucleotide.
To summarize the different approaches are:
- CAS9 and gRNA expressed by the same vector, each one under a specific promoter (All In One Vector)
- Two plasmids: one expressing gRNA and one expressing CAS9
- CAS9 plasmid + guide RNA (oligonucleotide RNA)
- In order to avoid Off Target CAS9 can be transfected as mRNA and guide RNA as oligonucleotide, later transcribed by In Vitro Transcription (IVT)
- Purified CAS9 protein can be combined with guideRNA to form a ribonucleoprotein complex (RNP) to be delivered to cells for rapid and highly efficient genome editing.
PRODUCTS
- ALL IN ONE SYSTEMS
- All in One Vector
- Pre-Designed kit
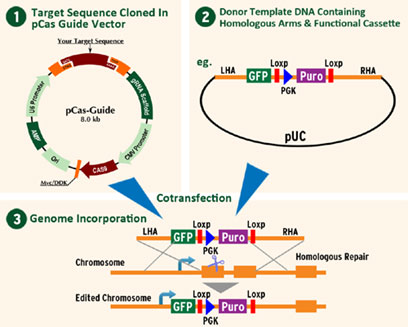
- Complete kit for gene knockout via CRISPR (targeted sites around the 5’ end of the ORF)
- 2 guide RNA vectors in pCas-Guide to ensure an efficient cleavage
- Donor vector with predesigned homologous arms
- Knockin GFP-Puro for selection
- pCas-Guide-scramble is also provided as a negative control
KN2.0 non-homology mediated CRISPR gene knockout kits
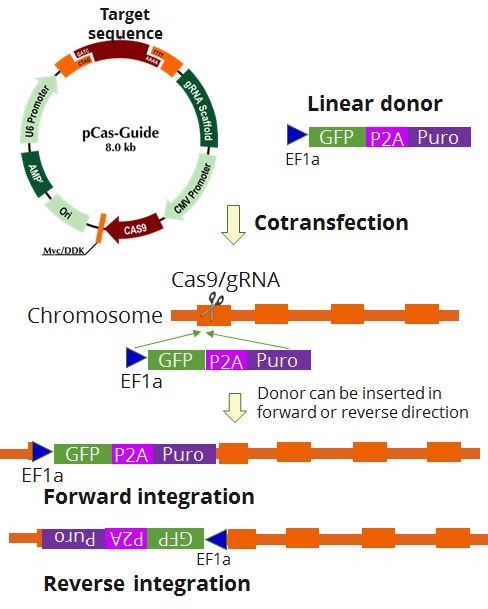
Gene knockout is based on non-homology-mediated repair mechanism. After gRNA targeted double stranded DNA cleavage, the linear donor DNA containing a selection cassette will be integrated at the gRNA cutting site at forward or reverse direction. Our testing data using KN2.0 kits showed that the vast majority gene knockout is bioallelic, one allele contains donor integration, the other allele has indels (insertion and deletion), which might change protein coding or cause premature stop.
Features
- Non-homology mediated gene knockout
- 2 guide RNA vectors in pCas-Guide
- Ready-to-use gene knockout kit
- Donor cassette, EF1a-GFP-P2A-Puro
- Highly efficient gene knockout kit
Scheme of gene knockout of KN2.0:
2. RNA OLIGONUCLEOTIDES
- CRISPR crRNA
- CRISPR tracrRNA
- S.p. CAS9 Nuclease 3NLS
- CRISPR-Cas9 control kits
- CRISPR positive and negative controls
- Cas9 expression plasmid
- Control PCR primer mixes
- Nuclease-Free Duplex Buffer
For Homology Directed Repair (HDR) you can combine:
RELATED PRODUCTS:
-
CRISPR/CAS9 TRANSFECTION:
Plasmid DNA (pDNA)+gRNA and RNP Delivery:
TransIT- X2®Dynamic Delivery System
mRNA + gRNA:
TransIT®-mRNA
-
ANTIBODIES
CAS9 Antibodies